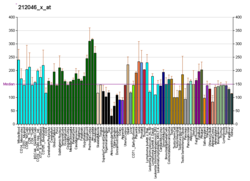
| MAPK3 |
|---|
|
| Estruturas disponíveis |
|---|
| PDB | Pesquisa Human UniProt: PDBe RCSB |
|---|
| Lista de códigos id do PDB |
|---|
2ZOQ, 4QTB |
|
|
| Identificadores |
|---|
| Nomes alternativos | MAPK3, p44MAPK, ERT2, ERK1 |
|---|
| IDs externos | OMIM: 601795 HomoloGene: 55682 GeneCards: MAPK3 |
|---|
|
| Targeted by Drug |
|---|
| ravoxertinib, ulixertinib[1] |
| Ontologia genética |
|---|
| Função molecular | • phosphatase binding
• ATP binding
• protein kinase activity
• MAP kinase activity
• transferase activity
• phosphotyrosine residue binding
• scaffold protein binding
• GO:0001948, GO:0016582 ligação a proteínas plasmáticas
• nucleotide binding
• kinase activity
• protein serine/threonine kinase activity
• MAP kinase kinase activity
• identical protein binding
|
|---|
| Componente celular | • citosol
• envelope nuclear
• focal adhesion
• mitocôndria
• citoesqueleto
• núcleo celular
• late endosome
• complexo de Golgi
• early endosome
• pseudópode
• cariolinfa
• citoplasma
• membrana plasmática
• caveola
• membrane
• GO:0009327 complexo macromolecular
|
|---|
| Processo biológico | • caveolin-mediated endocytosis
• positive regulation of protein phosphorylation
• positive regulation of telomere capping
• response to exogenous dsRNA
• cardiac neural crest cell development involved in heart development
• positive regulation of xenophagy
• positive regulation of translation
• cellular response to DNA damage stimulus
• platelet activation
• Fc-epsilon receptor signaling pathway
• protein phosphorylation
• cellular response to mechanical stimulus
• face development
• GO:0033129 positive regulation of histone modification
• regulation of DNA-binding transcription factor activity
• DNA damage induced protein phosphorylation
• positive regulation of ERK1 and ERK2 cascade
• animal organ morphogenesis
• ciclo celular
• GO:0097285 apoptose
• Fc-gamma receptor signaling pathway involved in phagocytosis
• thymus development
• ERK1 and ERK2 cascade
• negative regulation of apolipoprotein binding
• transcription, DNA-templated
• cartilage development
• GO:0022415 viral process
• response to toxic substance
• regulation of stress-activated MAPK cascade
• fosforilação
• outer ear morphogenesis
• BMP signaling pathway
• response to lipopolysaccharide
• thyroid gland development
• response to epidermal growth factor
• positive regulation of MAP kinase activity
• positive regulation of telomerase activity
• peptidyl-tyrosine autophosphorylation
• nocicepção
• positive regulation of cyclase activity
• trachea formation
• lipopolysaccharide-mediated signaling pathway
• GO:0007243 intracellular signal transduction
• lung morphogenesis
• neural crest cell development
• transcription initiation from RNA polymerase I promoter
• regulation of early endosome to late endosome transport
• positive regulation of telomere maintenance via telomerase
• MAPK cascade
• positive regulation of histone acetylation
• Guia do axônio
• interleukin-1-mediated signaling pathway
• fibroblast growth factor receptor signaling pathway
• GO:0035404 peptidyl-serine phosphorylation
• regulation of cellular response to heat
• Bergmann glial cell differentiation
• regulation of Golgi inheritance
• arachidonic acid metabolic process
• regulation of cytoskeleton organization
• GO:0003257, GO:0010735, GO:1901228, GO:1900622, GO:1904488 positive regulation of transcription by RNA polymerase II
• regulation of phosphatidylinositol 3-kinase signaling
• regulation of ossification
• GO:1901313 positive regulation of gene expression
• positive regulation of macrophage chemotaxis
• cellular response to amino acid starvation
• GO:0072353 cellular response to reactive oxygen species
• stress-activated MAPK cascade
• cellular response to cadmium ion
• cellular response to dopamine
• positive regulation of metallopeptidase activity
• regulation of cellular pH
• GO:0034622 protein-containing complex assembly
• cellular response to tumor necrosis factor
• regulação genética
• cellular response to organic substance
• GO:0010260 envelhecimento
• decidualization
|
|---|
| Sources:Amigo / QuickGO |
|
|
| Ortólogos |
|---|
| Espécie | Humano | Rato |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | |
|---|
NM_001040056
NM_001109891
NM_002746 |
| |
|---|
| RefSeq (proteína) | |
|---|
NP_001035145
NP_001103361
NP_002737 |
| |
|---|
| Localização (UCSC) | n/a | n/a |
|---|
| Pesquisa PubMed | [2] | n/a |
|---|
|
| Wikidata |
|
ERT2, também conhecida como p44MAPK, MAPK3 e ERK1, é uma enzima que em humanos é codificada pelo gene MAPK3.[3]
Ela é um membro da superfamília de proteína quinase, família de proteína cinase CMGC Ser/Thr, subfamília de MAP quinase. ERT2 é um apelido para o gene, proteína quinase 3 (ativada por mitógeno), em seres humanos.[4][5] Os pesquisadores fundiram a enzima a ERT2, em seguida ativaram o complexo com um análogo de tamoxifeno, 4-hidroxitamoxifeno, no caso de Cas9, a fim de colocar os controles no sistema CRISPR/Cas9.[6].
Interações
MAPK3 demonstrou interagir com:
- DUSP3,[7]
- DUSP6[8]
- GTF2I,[9]
- HDAC4,[10]
- MAP2K1,[11][12][13][14][15]
- MAP2K2,[11][12][15]
- PTPN7,[16][17][18]
- RPS6KA2,[19][20] and
- SPIB.[21]
Leitura adicional
- Peruzzi F, Gordon J, Darbinian N, Amini S (2002). «Tat-induced deregulation of neuronal differentiation and survival by nerve growth factor pathway». J. Neurovirol. 8 Suppl 2 (2): 91–6. PMID 12491158. doi:10.1080/13550280290167885
- Meloche S, Pouysségur J (2007). «The ERK1/2 mitogen-activated protein kinase pathway as a master regulator of the G1- to S-phase transition». Oncogene. 26 (22): 3227–39. PMID 17496918. doi:10.1038/sj.onc.1210414
- Ruscica M, Dozio E, Motta M, Magni P (2007), «Modulatory Actions of Neuropeptide y on Prostate Cancer Growth: Role of MAP Kinase/ERK 1/2 Activatio», Modulatory actions of neuropeptide Y on prostate cancer growth: role of MAP kinase/ERK 1/2 activation, ISBN 978-0-387-69114-5, Advances In Experimental Medicine And Biology, 604, pp. 96–100, PMID 17695723, doi:10.1007/978-0-387-69116-9_7
Referências
- ↑ «Drogas que interagem fisicamente com mitogen-activated protein kinase 3 ver/editar referências no wikidata»
- ↑ «Human PubMed Reference:»
- ↑ Garcı́a, F.; Zalba, G.; Páez, G.; Encı́o, I.; de Miguel, C. (15 de maio de 1998). «Molecular Cloning and Characterization of the Human p44 Mitogen-Activated Protein Kinase Gene». Genomics (em inglês) (1): 69–78. ISSN 0888-7543. doi:10.1006/geno.1998.5315. Consultado em 30 de junho de 2021
- ↑ Anti-ERT2 Antibody Products from Bio-Rad Antibodies publicado por "Biocompare"
- ↑ From Gene Targeting to Genome Editing: Transgenic animals applications and beyond por Mauricio Rocha-Martins et al, publicado pela An. Acad. Bras. Ciênc. vol.87 no.2 supl.0 Rio de Janeiro Aug. 2015 ISSN 1678-2690
- ↑ Toggling CRISPR Activity with a Chemical Switch Researchers design a Cas9 enzyme that cuts DNA only in the presence of particular drug. por Kerry Grens (2016)
- ↑ Todd JL, Tanner KG, Denu JM (maio de 1999). «Extracellular regulated kinases (ERK) 1 and ERK2 are authentic substrates for the dual-specificity protein-tyrosine phosphatase VHR. A novel role in down-regulating the ERK pathway». J. Biol. Chem. 274 (19): 13271–80. PMID 10224087. doi:10.1074/jbc.274.19.13271
- ↑ Muda M, Theodosiou A, Gillieron C, Smith A, Chabert C, Camps M, Boschert U, Rodrigues N, Davies K, Ashworth A, Arkinstall S (abril de 1998). «The mitogen-activated protein kinase phosphatase-3 N-terminal noncatalytic region is responsible for tight substrate binding and enzymatic specificity». J. Biol. Chem. 273 (15): 9323–9. PMID 9535927. doi:10.1074/jbc.273.15.9323
- ↑ Kim DW, Cochran BH (fevereiro de 2000). «Extracellular signal-regulated kinase binds to TFII-I and regulates its activation of the c-fos promoter». Mol. Cell. Biol. 20 (4): 1140–8. PMC 85232
 . PMID 10648599. doi:10.1128/mcb.20.4.1140-1148.2000
. PMID 10648599. doi:10.1128/mcb.20.4.1140-1148.2000 - ↑ Zhou X, Richon VM, Wang AH, Yang XJ, Rifkind RA, Marks PA (dezembro de 2000). «Histone deacetylase 4 associates with extracellular signal-regulated kinases 1 and 2, and its cellular localization is regulated by oncogenic Ras». Proc. Natl. Acad. Sci. U.S.A. 97 (26): 14329–33. PMC 18918
 . PMID 11114188. doi:10.1073/pnas.250494697
. PMID 11114188. doi:10.1073/pnas.250494697 - ↑ a b Marti A, Luo Z, Cunningham C, Ohta Y, Hartwig J, Stossel TP, Kyriakis JM, Avruch J (janeiro de 1997). «Actin-binding protein-280 binds the stress-activated protein kinase (SAPK) activator SEK-1 and is required for tumor necrosis factor-alpha activation of SAPK in melanoma cells». J. Biol. Chem. 272 (5): 2620–8. PMID 9006895. doi:10.1074/jbc.272.5.2620
- ↑ a b Butch ER, Guan KL (fevereiro de 1996). «Characterization of ERK1 activation site mutants and the effect on recognition by MEK1 and MEK2». J. Biol. Chem. 271 (8): 4230–5. PMID 8626767. doi:10.1074/jbc.271.8.4230
- ↑ Elion EA (setembro de 1998). «Routing MAP kinase cascades». Science. 281 (5383): 1625–6. PMID 9767029. doi:10.1126/science.281.5383.1625
- ↑ Yung Y, Yao Z, Hanoch T, Seger R (maio de 2000). «ERK1b, a 46-kDa ERK isoform that is differentially regulated by MEK». J. Biol. Chem. 275 (21): 15799–808. PMID 10748187. doi:10.1074/jbc.M910060199
- ↑ a b Zheng CF, Guan KL (novembro de 1993). «Properties of MEKs, the kinases that phosphorylate and activate the extracellular signal-regulated kinases». J. Biol. Chem. 268 (32): 23933–9. PMID 8226933
- ↑ Pettiford SM, Herbst R (fevereiro de 2000). «The MAP-kinase ERK2 is a specific substrate of the protein tyrosine phosphatase HePTP». Oncogene. 19 (7): 858–69. PMID 10702794. doi:10.1038/sj.onc.1203408
- ↑ Saxena M, Williams S, Taskén K, Mustelin T (setembro de 1999). «Crosstalk between cAMP-dependent kinase and MAP kinase through a protein tyrosine phosphatase». Nat. Cell Biol. 1 (5): 305–11. PMID 10559944. doi:10.1038/13024
- ↑ Saxena M, Williams S, Brockdorff J, Gilman J, Mustelin T (abril de 1999). «Inhibition of T cell signaling by mitogen-activated protein kinase-targeted hematopoietic tyrosine phosphatase (HePTP)». J. Biol. Chem. 274 (17): 11693–700. PMID 10206983. doi:10.1074/jbc.274.17.11693
- ↑ Roux PP, Richards SA, Blenis J (julho de 2003). «Phosphorylation of p90 ribosomal S6 kinase (RSK) regulates extracellular signal-regulated kinase docking and RSK activity». Mol. Cell. Biol. 23 (14): 4796–804. PMC 162206
 . PMID 12832467. doi:10.1128/mcb.23.14.4796-4804.2003
. PMID 12832467. doi:10.1128/mcb.23.14.4796-4804.2003 - ↑ Zhao Y, Bjorbaek C, Moller DE (novembro de 1996). «Regulation and interaction of pp90(rsk) isoforms with mitogen-activated protein kinases». J. Biol. Chem. 271 (47): 29773–9. PMID 8939914. doi:10.1074/jbc.271.47.29773
- ↑ Mao C, Ray-Gallet D, Tavitian A, Moreau-Gachelin F (fevereiro de 1996). «Differential phosphorylations of Spi-B and Spi-1 transcription factors». Oncogene. 12 (4): 863–73. PMID 8632909
Ligações externas
- localização do gene humano no navegador do genoma da UCSC.
- detalhes do gene humano no navegador do genoma da UCSC.
 Portal da genética
Portal da genética Portal da bioquímica
Portal da bioquímica
 . PMID 10648599. doi:10.1128/mcb.20.4.1140-1148.2000
. PMID 10648599. doi:10.1128/mcb.20.4.1140-1148.2000  . PMID 11114188. doi:10.1073/pnas.250494697
. PMID 11114188. doi:10.1073/pnas.250494697  . PMID 12832467. doi:10.1128/mcb.23.14.4796-4804.2003
. PMID 12832467. doi:10.1128/mcb.23.14.4796-4804.2003  Portal da genética
Portal da genética Portal da bioquímica
Portal da bioquímica