Crinivirus
| Crinivirus | |
|---|---|
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Kitrinoviricota |
| Class: | Alsuviricetes |
| Order: | Martellivirales |
| Family: | Closteroviridae |
| Genus: | Crinivirus |
| Species | |
| See text | |
| 3'-terminal pseudoknot in SPCSV | |
|---|---|
 Predicted secondary structure of the 3'-terminal pseudoknot in SPCSV | |
| Identifiers | |
| Rfam | RF01091 |
| Other data | |
| RNA type | Cis-reg |
| Domain(s) | Crinivirus |
| PDB structures | PDBe |
Crinivirus, formerly the lettuce infectious yellows virus group, is a genus of viruses, in the family Closteroviridae.[1] They are linear, single-stranded positive sense RNA viruses (and are therefore group IV).[2] There are 14 species in this genus.[1] Diseases associated with this genus include: yellowing and necrosis, particularly affecting the phloem.[1][3]
Examples of species whose entire genomes have been sequenced that are currently classified into the genus include the Sweet potato chlorotic stunt virus (SPCSV) and the Lettuce infectious yellows virus (LIYV).[4][5]
Genetics
The viruses of this genus have segmented, bipartite genomes that add up to 7,500–19,500 nucleotides in length. Their genomes also code for proteins that do not form part of the virion particles as well as structural proteins. The Universal Virus Database describes that their genome sequences near their 3'-ends are capable of hairpin-loop formation and also believe that their 5'-ends may have methylated caps.[5] Each of the viral RNA molecules contains four hair-pin structures and a pseudoknot in the 3'UTR. The pseudoknot is unusual in that it contains a small stem-loop structure inside loop L1.[6] In the related genus Closterovirus, these secondary structures have been found to be important in viral RNA replication.[7]
Structure
Viruses in the genus Crinivirus are non-enveloped, with bipartite filamentous geometries. The diameter is around 10-13 nm, with a length of 700-900 nm. Genomes are linear and bipartite, around 17.6kb in length.[1][3]
| Genus | Structure | Symmetry | Capsid | Genomic arrangement | Genomic segmentation |
|---|---|---|---|---|---|
| Crinivirus | Filamentous | Non-enveloped | Linear | Monopartite, bipartite, or tripartite |
Life cycle
Viral replication is cytoplasmic. Entry into the host cell is achieved by penetration into the host cell. Replication follows the positive stranded RNA virus replication model. Positive stranded RNA virus transcription is the method of transcription. The virus exits the host cell by tubule-guided viral movement. Plants serve as the natural host. The virus is transmitted via a vector (bemisia tabaci). Transmission route is mechanical.[1][3]
| Genus | Host details | Tissue tropism | Entry details | Release details | Replication site | Assembly site | Transmission |
|---|---|---|---|---|---|---|---|
| Crinivirus | Plants | None | Viral movement; mechanical inoculation | Viral movement | Cytoplasm | Cytoplasm | Mechanical inoculation: insects (whitefly) |
Taxonomy
The following species are assigned to the genus:[8]
- Abutilon yellows virus
- Bean yellow disorder virus
- Beet pseudoyellows virus
- Blackberry yellow vein-associated virus
- Cucurbit yellow stunting disorder virus
- Diodia vein chlorosis virus
- Lettuce chlorosis virus
- Lettuce infectious yellows virus
- Potato yellow vein virus
- Strawberry pallidosis-associated virus
- Sweet potato chlorotic stunt virus
- Tetterwort vein chlorosis virus
- Tomato chlorosis virus
- Tomato infectious chlorosis virus
References
- ^ a b c d e "ICTV Report Closteroviridae".
- ^ ICTVdB Management (2006). 00.017.0.02. Crinivirus. In: ICTVdB—The Universal Virus Database, version 4. Büchen-Osmond, C. (Ed), Columbia University, New York, USA
- ^ a b c "Viral Zone". ExPASy. Retrieved 15 June 2015.
- ^ Journal of Virology. 2002 September; 76(18): 9260–9270
- ^ a b ICTVdB Management (2006)
- ^ Livieratos IC, Eliasco E, Müller G, et al. (July 2004). "Analysis of the RNA of Potato yellow vein virus: evidence for a tripartite genome and conserved 3'-terminal structures among members of the genus Crinivirus". J. Gen. Virol. 85 (Pt 7): 2065–75. doi:10.1099/vir.0.79910-0. hdl:1887/3629824. PMID 15218192.
- ^ Satyanarayana T, Gowda S, Ayllón MA, Albiach-Martí MR, Dawson WO (August 2002). "Mutational analysis of the replication signals in the 3'-nontranslated region of citrus tristeza virus". Virology. 300 (1): 140–52. doi:10.1006/viro.2002.1550. PMID 12202214.
- ^ "Virus Taxonomy: 2020 Release". International Committee on Taxonomy of Viruses (ICTV). March 2021. Retrieved 14 May 2021.
External links

- ICTV Report: Closteroviridae
- Viralzone: Crinivirus
- Rfam entry for 3'-terminal pseudoknot in SPCSV
- Rfam entry for 3'-terminal pseudoknot of CuYV/BPYV
- Rfam entry for 3'-terminal pseudoknot in PYVV
- v
- t
- e
-
 3'-terminal pseudoknot in SPCSV: Predicted secondary structure taken from the Rfam database. Family RF01091.
3'-terminal pseudoknot in SPCSV: Predicted secondary structure taken from the Rfam database. Family RF01091. -
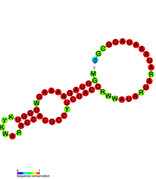 3'-terminal pseudoknot in PYVV: Predicted secondary structure taken from the Rfam database. Family RF01078.
3'-terminal pseudoknot in PYVV: Predicted secondary structure taken from the Rfam database. Family RF01078. -
 3'-terminal pseudoknot of CuYV/BPYV: Predicted secondary structure taken from the Rfam database. Family RF01095.
3'-terminal pseudoknot of CuYV/BPYV: Predicted secondary structure taken from the Rfam database. Family RF01095.













